


 النبات
النبات
 الحيوان
الحيوان
 الأحياء المجهرية
الأحياء المجهرية
 علم الأمراض
علم الأمراض
 التقانة الإحيائية
التقانة الإحيائية
 التقنية الحيوية المكروبية
التقنية الحيوية المكروبية
 التقنية الحياتية النانوية
التقنية الحياتية النانوية
 علم الأجنة
علم الأجنة
 الأحياء الجزيئي
الأحياء الجزيئي
 علم وظائف الأعضاء
علم وظائف الأعضاء
 الغدد
الغدد
 المضادات الحيوية
المضادات الحيوية|
Read More
Date: 17-6-2021
Date: 28-12-2015
Date: 22-12-2015
|
Chemotaxis
Chemotaxis is the movement of a motile cell in response to chemical changes in the environment, and it implies a directional sense. In most cases where bacteria migrate up concentration gradients of attractants, direction is not sensed directly. The small size of bacteria would make spatial comparisons of concentration unreliable. Instead, bacteria usually sense temporal changes in concentration by using chemoreceptors, transducer proteins that bind or release attractants and repellents. This temporal measurement allows the cells to alter motion of their flagella briefly so that a “biased random walk” brings the bacteria to the right destination (1, 2) (Fig. 1). Eukaryotic cells carry out chemotaxis by an unrelated mechanism, and they are much larger and probably do sense direction.
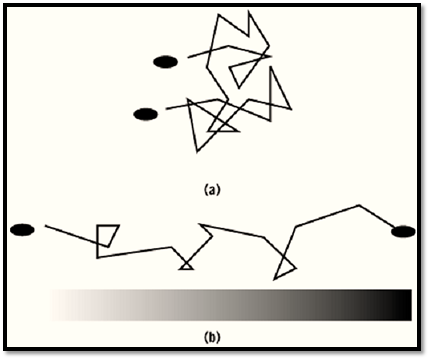
Figure 1. Motion of peritrichous bacteria. (a) Random walk in an isotropic medium. (b) Biased random walk in a gradient of an attractant or repellent.
1. Bacteria
There are three main types of motile bacteria: 1) the peritrichous (flagella over the surface), 2) the polar (flagella at one or both poles), and 3) the gliding bacteria, which lack flagella.
The peritrichous bacteria, such as Escherichia coli and Bacillus subtilis, normally swim smoothly by rotating their flagella counterclockwise (looking along the flagella toward the cell body) and tumble by rotating their flagella clockwise, which discoordinates the bundle of flagella. Chemotaxis in E. coli occurs when the cell happens to move toward higher attractant concentrations. It decreases the probability of tumbling and, thus, increases the tendency to travel up the attractant gradient (1).
This decreased probability of tumbling results (Fig. 2) because the attractant molecules bind to methylated chemoreceptors to cause a brief inactivation of the cytoplasmic CheA kinase. Consequently, there is less phosphorylated CheA (CheA-P) and less CheY-P (2). CheY-P dissociates from the switch that controls flagellar rotation, so that smooth swimming ensues. The enhanced smooth swimming is short-lived because CheZ hydrolyzes the phosphoryl group from CheY, and the CheR methyltransferase increases the degree of methylation of the chemoreceptor and restores the prestimulus activity of CheA (3). Thus, there is an inverse relation between the levels of CheY-P and termination of a smooth swimming event.
Logically, it seems plausible that bacteria moving by chance toward lower attractant concentrations would show an increased tendency to tumble and, hence, to begin heading in another direction. In the case of E. coli, however, there is no demonstrable increased tendency for these bacteria to tumble compared with bacteria in an isotropic medium (1). Interestingly, CheY-P cannot affect rotational direction of the flagella by itself. Fumarate needs to be released or synthesized for switching to occur (4) and also promotes tumbling (5). The connection between CheY-P and fumarate in controlling direction of flagellar rotation has not been elucidated.
The role of CheY-P is less clear in the case of polar bacteria, such as Caulobacter crescentus and Pseudomonas aeruginosa. When these bacteria move by chance toward lower concentrations of attractant, they reverse motion by rotating their flagella in the opposite direction, to head back up the gradient. If repellent is added, however, they show a series of reversals of motion. This result implies that CheY-P does not govern the direction of rotation of the flagella but instead causes repeated reversals of behavior. Such an event might ensue if binding of CheY-P to the switch reduces the energy of activation of switching the flagellar rotational direction. In the absence of binding, the energy of activation is presumed to be so high that reversals are very infrequent. In the case of Halobacterium salinarium, negative stimuli cause significant increases in the concentrations of free fumarate in the cytoplasm (4).
Other bacteria, like Sinorhizoboium meliloti, a Gram-negative bacterium phylogenetically rather distant from E. coli, rotate their flagella unidirectionally. In this case, the CheA/CheY system also exists with the complication that there are two species of CheY. CheY2-P is the major species that affects flagellar movement, and it acts by slowing flagellar rotation (6). Some bacteria, such as Rhodobacter sphaeroides, have multiple copies of many of the che genes (7).
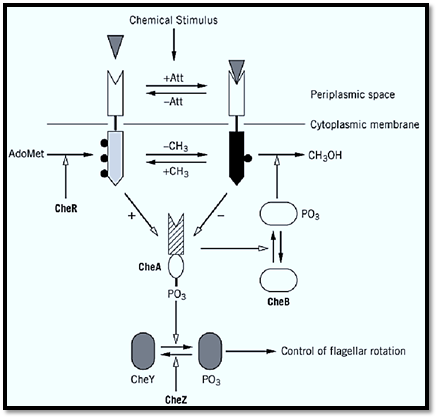
Figure 2. Diagram of the mechanism of chemotaxis in Escherichia coli. Open arrows indicate control of the designated reaction or signaling step. At the top, the attractant (Att) binds to its transmembrane chemoreceptor, which is methylated on Glu residues. The methyl groups are depicted as closed black circles. This binding decreases the phosphorylated form of the protein CheA (center), which decreases phosphorylation of the proteins CheY and CheB. Dephosphorylated CheB is much less active in removing the methyl groups from the chemoreceptor, so CheR restore them. Phosphorylated CheY dissociates from the switch that controls flagellar rotation, and smooth swimming ensues. CheZ hydrolyzes the phosphoryl group from CheY, ending the response to the attractant.
Thus, it appears that the basic system involving the two-component couple CheA and CheY is universal, but how it functions varies for different groups of organisms. The potential role for CheY-P is still less clear in the case of the gliding bacteria. One of them, Myxococcus xanthus, has two motility systems, the S (social) system powered by extrusion and retraction of pili from the ends and the A (adventurous) system, powered by a mechanism of unknown structure along the sides. The former coordinates cells and, unless the agar concentration is quite low, requires cell contact for movement. The frequency and pole at which pili activity occurs is governed by the Frz proteins, which are homologous to the chemotaxis proteins in flagellated cells. However, none of the genes encode a single-domain CheY (8, 9). One issue that has existed for many years has been whether the rotation of multiple flagella are coordinated or independent. The finding that bundles of flagella located 50 mm apart on Spirillum volutans were observed switching synchronously within 10 ms implies electrical coordination of direction of the flagellar rotation (1). In contrast, certain mutant strains of Salmonella typhimurium show random and uncorrelated switching of their flagellar direction when observed under partially de-energized conditions (10). E. coli filamentous cells switch their flagella asynchronously although biases of nearby, but not distant, motors were correlated (11). In H. salinarium, an archaeon, the rotational direction of flagellar bundles on each end is also uncoordinated, and cells were frequently observed to be immobilized because the bundles at both ends worked against each other. Thus, there is no general rule governing flagellar coordination. Indeed, the mechanism by which an electrical connection could work, as in S. volutans, is obscure.
2. Eukaryotes
Chemotaxis in eukaryotes occurs by an unrelated mechanism. The process is best known from studies in the aggregating slime mold Dictyostelium discoidium and in leucocytes; it is probably quite similar in both. In the slime mold, chemotaxis plays a role in feeding and in the aggregation that precedes differentiation and sporulation. Chemotaxis in Dictyostilium involves apparent polarization of the cell to form pseudopodia or lamellipodia at the leading edge at the highest concentration of attractant molecules. Polarization and formation of pseudopodia are concerted events regulated by G protein-coupled/serpentine cyclic AMP (cAMP) receptors (cARs) along the surface of the organism. During aggregation, cAMP is produced and binds to a cAR, which activates a heterotrimeric G-protein. Guanylate cyclase is activated by operation of this G-protein, thereby producing cGMP. cGMP activates a cGMP-dependent protein kinase, which regulates myosin II heavy and light chain kinases and may also help regulate the actin cytoskeleton. Phosphorylation of the myosin heavy chain causes it to depolymerize so that pseudopod extension can only occur locally. Posteriorly, contraction occurs and pseudopod formation is blocked. At the same time cAMP binding to its receptor leads to activation of phosphatidylinositol-3 kinase. The product, phosphoinositol-3,4,5-triphosphate, attracts PH (pleckstrin homology) proteins, including the kinase Akt/PKB, to the localized region where the local amount of cAMP-bound receptor is highest. Another PH domain protein is CRAC. It is required for receptor activation of adenylyl cyclase, which then propagates the cAMP signal to neighboring cells. This membrane localization lasts only 5 to 8 s; however, that is long enough to produce an “activation domain” as a focus for multiple pathways needed for chemotaxis, pseudopod extension, and cell polarization. F-actin is assembled primarily there. Actin polymerization is activated by formation of an Arp2/3 protein complex. Nucleation is induced by Arp2/3 interaction with WASp and Scar adaptor proteins. These proteins provide the structural linkage to the activated cAR through the activated form of Rac and Cdc42, which are regulated by the G protein Gbg. Pseudopod formation also requires actin crosslinking and filament growth regulated by actin binding proteins (12, 13).
Binding of cAMP to the receptors transiently activates both adenylyl cyclase and ERK2 MAP kinase. A rise in internal cAMP activates protein kinase A such that it inhibits ERK2 and leads to loss of ligand binding by the receptor. ERK2 also phosphorylates the cAMP diesterase REG A that reduces the internal concentration of cAMP. A secreted membrane-associated phosphodiesterase reduces the external cAMP concentrations, and the cells become sensitive again. These features account for spontaneous oscillations of external cAMP production of about 6 min that permit chemotactic migration of cells toward centers during the aggregation process (14).
References
1. H. C. Berg (1975) Annu. Rev. Biophys. Bioeng. 4, 119–136.
2. J. B. Stock and M. G. Surette (1996) In Escherichia coli and Salmonella, Cellular and Molecular Biology, 2 ed. (F. C. Neidhardt, ed.), ASM Press, Washington, DC, Vol. I, pp. 1103–1129.
3. J. S. Parkinson (1993) Cell 73, 857–871.
4. R. Barak and M. Eisenbach (1996) Curr. Top. Cell. Reg. 34, 137–158.
5. K. Prasad, S. R. Caplan, and M. Eisenbach (1998) J. Mol. Biol. 280, 821–828.
6. V. Sourjik and R. Schmitt (1996) Mol. Microbiol. 22, 427–436.
7. J. P. Armitage Adv. Microbiol. Physiol. 41, 229–289.
8. H. Sun, D. R. Zusman, and W. Shi (2000) Curr. Biol. 10, 1143–1146.
9. M. J. Ward and D. R. Zusman (1997) Mol. Microbiol. 24, 885–893.
10. R. M. Macnab and D. P. Han (1983) Cell 32, 109–117.
11. A. Ishihara, J. E. Segall, S. M. Block, and H. C. Berg (1983) J. Bacteriol. 155, 228–237.
12. R. A. Firtel and R. Meili (2000) Curr. Opin. Genet. Develop. 10, 421–427.
13. R. A. Firtel and C. Y. Chung (2000) BioEssays 22, 603–615.
14. M. T. Laub and W. F. Loomis (1998) Mol. Biol. Cell 12, 3521–3532.

|
|
|
|
علامات بسيطة في جسدك قد تنذر بمرض "قاتل"
|
|
|
|
|
|
|
أول صور ثلاثية الأبعاد للغدة الزعترية البشرية
|
|
|
|
|
|
|
مدرسة دار العلم.. صرح علميّ متميز في كربلاء لنشر علوم أهل البيت (عليهم السلام)
|
|
|