


 النبات
النبات
 الحيوان
الحيوان
 الأحياء المجهرية
الأحياء المجهرية
 علم الأمراض
علم الأمراض
 التقانة الإحيائية
التقانة الإحيائية
 التقنية الحيوية المكروبية
التقنية الحيوية المكروبية
 التقنية الحياتية النانوية
التقنية الحياتية النانوية
 علم الأجنة
علم الأجنة
 الأحياء الجزيئي
الأحياء الجزيئي
 علم وظائف الأعضاء
علم وظائف الأعضاء
 الغدد
الغدد
 المضادات الحيوية
المضادات الحيوية|
Read More
Date: 18-11-2020
Date: 27-2-2021
Date: 5-5-2021
|
The Clamp Controls Association of Core Enzyme with DNA
KEY CONCEPTS
- The core on the leading strand is processive because its clamp keeps it on the DNA.
- The clamp associated with the core on the lagging strand dissociates at the end of each Okazaki fragment and reassembles for the next fragment.
- The helicase DnaB is responsible for interacting with the primase DnaG to initiate each Okazaki fragment.
The β2 -ring dimer makes the holoenzyme highly processive. β is strongly bound to DNA but can slide along a duplex molecule. The crystal structure of β shows that it forms a ring-shaped dimer. The model in FIGURE 1 shows the β2 ring in relationship to a DNA double helix. The ring has an external diameter of 80 Å and an internal cavity of 35 Å, almost twice the diameter of the DNA double helix (20 Å). The space between the protein ring and the DNA is filled by water. Each of the β subunits has three globular domains with similar organization (although their sequences are different). As a result, the dimer has sixfold symmetry that is reflected in 12 α-helices that line the inside of the ring.

FIGURE 1 The subunit of DNA polymerase III holoenzyme consists of a head-to-tail dimer (the two subunits are shown in red and orange) that forms a ring completely surrounding a DNA duplex (shown in the center).
Reprinted from: Kong, X. P., et al. 1992. “Three-dimensional structure of the β.” Cell 69:425–437, with permission from Elsevier (http://www.sciencedirect.com/science/journal/00928674). Photo courtesy of John Kuriyan, University of California, Berkeley.
The β2 -ring dimer surrounds the duplex, providing the “sliding clamp” that allows the holoenzyme to slide along DNA. The structure explains the high processivity—the enzyme can transiently dissociate but cannot fall off and diffuse away. The α-helices on the inside have some positive charges that might interact with the DNA via the intermediate water molecules. The protein clamp does not directly contact the DNA, and, as a result, it might be able to “ice skate” along the DNA, making and breaking contacts via the water molecules.
How does the clamp get onto the DNA? The clamp is a circle of subunits surrounding DNA; thus, its assembly or removal requires the use of an energy-dependent process by the clamp loader. The γ clamp loader is a pentameric circular structure that binds an open form of the β2 ring preparatory to loading it onto DNA. In effect, the ring is opened at one of the interfaces between the two β2 subunits by the δ subunit of the clamp loader. The binding of δ to the ring destabilizes and opens it, facilitated by ATP. The role of ATP is not clear, whether hydrolysis is used to open the β2 ring or for release of the clamp loader. The SSB proteins that coat the DNA are not passive, but rather are required to stimulate the process. The relationship between the β2 clamp and the γ clamp loader is a paradigm for similar systems used by DNA polymerases ranging from bacteriophages to animal cells. The clamp is a heteromer (possibly a dimer or trimer) that forms a ring around DNA with a set of 12 α-helices forming sixfold symmetry for the structure as a whole. The clamp loader has some subunits that hydrolyze ATP to provide energy for the reaction.
The basic principle that is established by the dimeric polymerase model is that, while one polymerase subunit synthesizes the leading strand continuously, the other cyclically initiates and terminates the Okazaki fragments of the lagging strand within a large, singlestranded loop formed by its template strand. FIGURE 2 draws a generic model for the operation of such a replicase. The replication fork is created by a helicase—which typically forms a hexameric ring—that translocates in the 5′–3′ direction on the template for the lagging strand. The helicase is connected to two DNA polymerase catalytic subunits, each of which is associated with a sliding clamp.
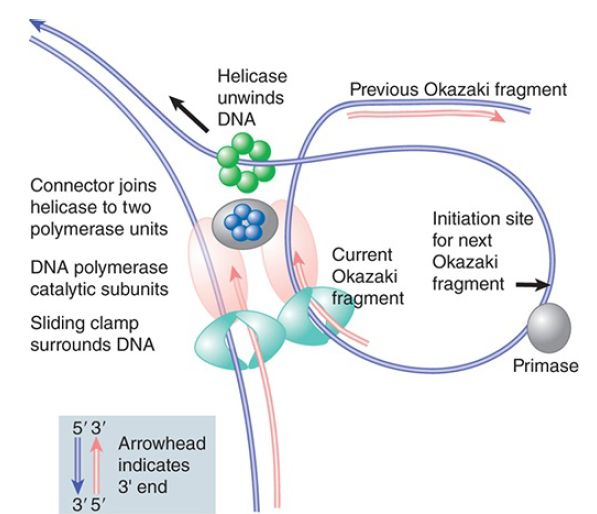
FIGURE 2. The helicase creating the replication fork is connected to two DNA polymerase catalytic subunits, each of which is held onto DNA by a sliding clamp. The polymerase that synthesizes the leading strand moves continuously. The polymerase that synthesizes the lagging strand dissociates at the end of an Okazaki fragment and then reassociates with a primer in the singlestranded template loop to synthesize the next fragment.
We can describe this model for DNA polymerase III in terms of the individual components of the enzyme complex, as illustrated in FIGURE 3. A catalytic core is associated with each template strand of DNA. The holoenzyme moves continuously along the template for the leading strand; the template for the lagging strand is “pulled through,” thus creating a loop in the DNA. DnaB creates the unwinding point and translocates along the DNA in the “forward” direction.
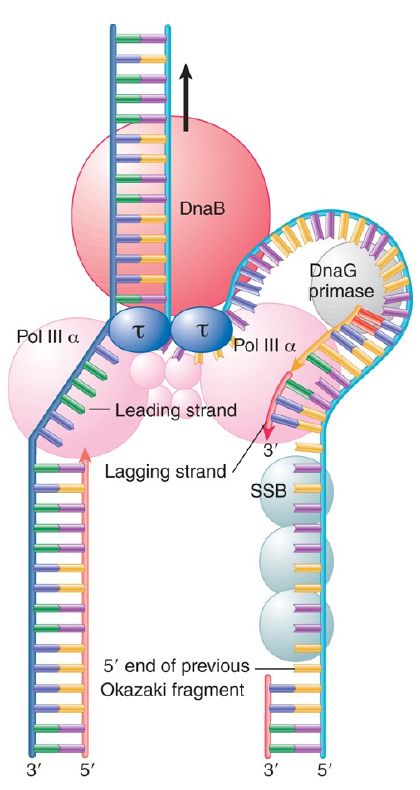
FIGURE 3. Each catalytic core of Pol III synthesizes a daughter strand. DnaB is responsible for forward movement at the replication fork. DnaB contacts the τ subunit(s) of the clamp loader. This establishes a direct connection between the helicase–primase complex and the catalytic cores. The link has two effects. One is to increase the speed of DNA synthesis by increasing the rate of movement by DNA polymerase core by 10-fold. The second is to prevent the leading strand polymerase from falling off, that is, to increase its processivity.
Synthesis of the leading strand creates a loop of single-stranded DNA that provides the template for lagging strand synthesis, and this loop becomes larger as the unwinding point advances. After initiation of an Okazaki fragment, the lagging strand core complex pulls the single-stranded template through the β2 clamp while synthesizing the new strand. The single-stranded template must extend for the length of at least one Okazaki fragment before the lagging polymerase completes one fragment and is ready to begin the next.
What happens when the Okazaki fragment is completed? All of the components of the replication apparatus function processively (i.e., they remain associated with the DNA), except for the primase and the β2 clamp. FIGURE 4 shows that the β2 clamp must be cracked open by the γ clamp loader when the synthesis of each fragment is completed, releasing the loop. We can think of the clamp loader here as a molecular wrench that is modulated by ATP.
The clamp loader causes the β2 clamp to alter its conformation to an unstable configuration, which then springs open. A new β2 clamp is then recruited by the clamp loader to initiate the next Okazaki fragment. The lagging strand polymerase transfers from one β2 clamp to the next in each cycle, without dissociating from the replicating complex.

FIGURE 4. Core polymerase and the clamp dissociate at completion of Okazaki fragment synthesis and reassociate at the beginning.
What is responsible for recognizing the sites for initiating synthesis of Okazaki fragments? In oriC replicons, the connection between priming and the replication fork is provided by the dual properties of DnaB: It is the helicase that propels the replication fork, and it interacts with the DnaG primase at an appropriate site. Following primer synthesis, the primase is released. The length of the priming RNA is limited to 8 to 14 bases. Apparently, DNA polymerase III is responsible for displacing the primase.

|
|
|
|
تفوقت في الاختبار على الجميع.. فاكهة "خارقة" في عالم التغذية
|
|
|
|
|
|
|
أمين عام أوبك: النفط الخام والغاز الطبيعي "هبة من الله"
|
|
|
|
|
|
|
قسم شؤون المعارف ينظم دورة عن آليات عمل الفهارس الفنية للموسوعات والكتب لملاكاته
|
|
|