


 النبات
النبات
 الحيوان
الحيوان
 الأحياء المجهرية
الأحياء المجهرية
 علم الأمراض
علم الأمراض
 التقانة الإحيائية
التقانة الإحيائية
 التقنية الحيوية المكروبية
التقنية الحيوية المكروبية
 التقنية الحياتية النانوية
التقنية الحياتية النانوية
 علم الأجنة
علم الأجنة
 الأحياء الجزيئي
الأحياء الجزيئي
 علم وظائف الأعضاء
علم وظائف الأعضاء
 الغدد
الغدد
 المضادات الحيوية
المضادات الحيوية|
Read More
Date: 27-4-2016
Date: 1-11-2015
Date: 29-5-2021
|
Achaete–Scute Complex
A central theme in developmental biology is to understand how the mechanisms that specify particular cell types integrate with those that govern where and when cell type-specific structures form during animal development. Over the better part of the past century, the development of the Drosophila nervous system has served as a model with which to unravel the interconnected processes of cell fate specification and pattern formation. The genes of the achaete–scute complex (AS-C) bridge these two processes. The AS-C is composed of four related genes found within ~90 kbp of DNA in the distal tip of the X-chromosome. The four genes—achaete (ac), scute (sc), lethal of scute (l'sc) and asense (ase)—all encode for basic helix-loop-helix transcriptional activator proteins and were initially identified by mutational analysis. Loss-of-function mutations in the AS-C remove sensory organs in the peripheral nervous system (PNS) and neural structures in the central nervous system (CNS), whereas gain-of-function mutations induce the formation of ectopic neural structures. The “proneural” ac, sc, and l'sc genes are expressed prior to neural precursor formation in precise patterns of cell clusters that forecast where neural precursors will form. During both embryonic and adult development, the primary patterning genes in Drosophila act through a large array of cis-acting regulatory regions found within the AS-C to create the stereotyped pattern of AS-C-expressing proneural cell clusters. Within each cluster (equivalence group), a cell communication process mediated by the Notch signaling pathway (lateral inhibition) restricts AS-C gene expression to a single cell. Within this cell, ac, sc and l'sc, either alone or in combination, are thought to activate a cassette of genes that directs this cell to acquire the neural precursor fate. All other cells within the cluster extinguish AS-C gene expression and are directed towards epidermal development. One target of the AS-C proneural genes in neural precursors is ase, which promotes the differentiation of neural structures. The AS-C thus links the process of cell fate specification to that of pattern formation: the AS-C genes receive and interpret global positional information through their regulatory regions and translate this information through their function into the formation (and differentiation) of a two-dimensional array of neural structures.
1. Genetic and Molecular Characterization of the Achaete–Scute Complex
The embryonic and adult Drosophila nervous systems are composed of a variety of different neural structures, each organized in reproducible but distinct patterns. For example, the neural precursors of the embryonic CNS form in orthogonal rows, whereas in the adult PNS large innervated bristles arise in an irregular, yet stereotyped pattern. Each bristle, or sensory organ, forms from a single neural precursor cell that occupies a fixed location in the developing fly. Cell migration is limited in the Drosophila PNS and CNS. Thus, the initial neural precursor pattern largely determines the later pattern of neural structures.
Genetic analyses identified a set of four genetic activities, designated the AS-C, located near the distal tip of the X-chromosome critical for the formation, patterning, and differentiation of neural structures (1). Loss-of-function mutations in any one of these genes remove neural structures, although the precise set of structures removed differs for each gene. For example, loss of l'sc activity primarily affects the CNS, whereas loss of ac or sc activity primarily results in PNS defects. Loss of ase function removes sensory organs along the wing blade but also causes differentiation defects in other sensory organs. Conversely, mis-expression of any AS-C gene promotes the formation of ectopic neural structures. These analyses suggested that the AS-C promotes the initial commitment of a cell to become a neural precursor and that their gene products are at least partially redundant. Detailed genetic mosaic analyses in the Drosophila wing imaginal disc (the larval structure that generates the wing and notum of the adult fly) showed that the activities of ac and sc define zones of neural competency from which neural precursors arise (2). These zones of neural competency are known as proneural cell clusters and are “equivalence groups” within which all cells can (although only one or a few will) become a neural precursor.
The genes ac, sc, and l'sc are each expressed in a complex and dynamic pattern of cell clusters, each of which quickly resolves to a single cell, the putative neural precursor (Fig. 1). The gene expression patterns of ac and sc are identical throughout neurogenesis and partially overlap with the expressionpattern of l'sc. On the other hand, asense is expressed exclusively in neural precursors and their progeny. Based on mutant phenotypes and expression patterns, ac, sc, and l'sc are called proneural genes whereas ase is called a neural precursor gene.
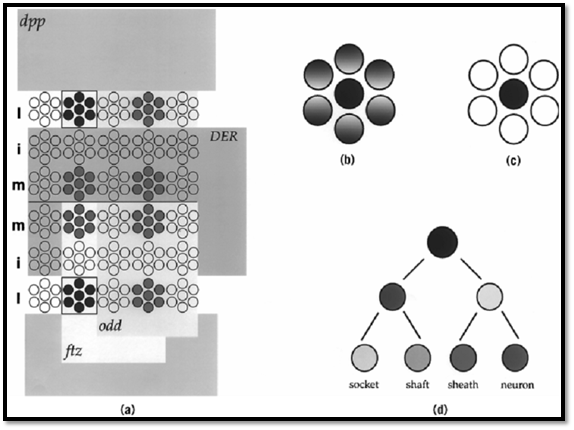
Figure 1. Patterning and specification of neural precursors. (a) Patterning of AS-C gene expression. Schematic representation of the ectoderm of a Drosophila embryo. The expression of the ac and sc genes in four proneural clusters per hemisegment is shown in red and blue. The activity of the early patterning genes ftz, odd, DER, and dpp subdivide the embryo and lead to the activation of ac/sc gene expression in reproducible quadrants. For example, the combined action of ftz activation and odd repression leads to ac/sc gene expression in domains that express ftz but not odd (yellow). Along the D/V axis, dpp (light blue) and DER (green) restrict ac/sc expression to the lateral column. Together, the action of these four genes delimit ac/sc gene expression within one proneural cluster, shown in blue. Note that other factors can overcome DER mediated repression in the medial column. (b) Singling out of a neural precursor. Within each proneural cluster, a single cell (in this example, the central cell) comes to express the AS-C proneural genes to the highest level, and this cell becomes the presumptive neural precursor (dark blue cell). (c) Resolution of cell fate within a proneural cluster. The presumptive neural precursor retains proneural gene expression and acts through the Notch signaling pathway to remove proneural gene expression in the other cells of the cluster (white). These cells are directed toward epidermal development. (d) Neural precursor lineage. Each neural precursor goes through an invariant cell lineage to produce a particular neural structure. The lineage of a sensory bristle of the PNS is shown. The neural precursor divides to yield two differently fated daughter cells, which in turn divide to produce a total of four cells. Each cell acquires a different fate, and together the four cells make up a functional sensory organ—an innervated bristle.
The molecular cloning of the AS-C in the 1980s identified all four genes as encoding putative transcriptional activator proteins that belong to the class B type of basic helix-loop-helix (bHLH) proteins. These four genes are contained within ~90 kbp of DNA, separated from one another by an average of ~30 kbp of DNA replete with cis-regulatory regions required to activate different members of the AS-C in complex and partially overlapping patterns (3). Transcriptional activation requires the formation of a heterodimer between A and B class bHLH proteins (4); these heterodimers recognize the core DNA sequence CANNTG, where N is not specified (an ’E’ box). An A-class bHLH protein with which AS-C gene products heterodimerize and activate transcription is the product of the daughterless (da) gene. Heterodimers of one AS-C gene product and Da mediate transcriptional activation of genes expressed throughout proneural clusters and in neural precursors. The da gene is expressed ubiquitously throughout Drosophila development, and AS-C genes are expressed in complex yet invariant patterns of cell clusters that predict where neural precursors form. Thus, the spatiotemporal specificity of where and when neural precursors develop is a function of the pattern of AS-C gene expression.
2. Cis- and Trans-Regulation of Proneural Gene Expression
Activation of the proneural genes of the AS-C in precise and reproducible patterns of proneural clusters is the first step in the formation of the stereotyped two-dimensional pattern of neural precursors in the Drosophila nervous system. The genetic regulatory mechanisms that dictate where and when AS-C expressing proneural clusters form are best understood for the development of the embryonic CNS (5). The anterior/posterior (A/P) and dorsal/ventral (D/V) register of AS-C-expressing cell clusters in the early embryo suggests that the segmentation genes, which segment the embryo along the A/P axis, and the dorsoventral genes, which specify pattern along the D/V axis, function to lay down directly the initial pattern of AS-C proneural clusters. A similar process mediated by wingless and decapentaplegic (dpp), as well as the genes of the iroquois complex (6), probably acts directly on the AS-C to create the pattern of AS-C-expressing cell clusters in adult structures.
During development of the embryonic Drosophila CNS, the AS-C proneural genes are expressed in an invariant orthogonal pattern of proneural cell clusters. For example, ac and sc are co-expressed in two rows of cell clusters in the medial and lateral—but not intermediate—columns of each segment. Prior to AS-C gene activation, the activities of different A/P and D/V patterning genes have subdivided the early embryo into an orthogonal pattern of squares, reminiscent of a checkerboard. Each square expresses a unique combination of these patterning genes, and the borders of the AS-C proneural clusters match precisely the limits of these squares (Fig. 1). This suggests a model whereby the combined activities of particular combinations of patterning genes within a square either does or does not activate a member of the AS-C. The composite pattern of AS-C-positive cell clusters would then result from the integration of the activities of each square.
Support for the checkerboard model comes from genetic analyses that assayed the expression of AS-C genes in embryos mutant for various known patterning genes. For example, the anterior border of every fourth transverse row of ac/sc-positive cell clusters coincides with the anterior edge of the expression of the fushi-tarazu (ftz) pair-rule gene, and the posterior border of these proneural clusters abuts the anterior border of the odd-skipped (odd) pair-rule gene (Fig. 1). In embryos mutant for ftz, this row of ac/sc-expressing proneural clusters disappears, whereas in embryos mutant for odd it expands posteriorly to fill the entire ftz domain. Thus, the combined action of ftz activation and odd repression defines the anterior and posterior boundaries respectively, of proneural clusters in this row. The D/V patterning genes control the mediolateral extent of AS-C proneural clusters. For example, the medial boundary of the lateral column of AS-C proneural clusters abuts the lateral border of the activity of the Drosophila epidermal growth factor (EGF) receptor (DER), and the lateral boundary of these clusters abuts the medial boundary of dpp activity (Fig. 1). In embryos mutant for DER, the lateral column of AS-C-expressing proneural clusters expands medially, whereas in dpp mutant embryos these clusters expand laterally. Thus, DER and dpp together restrict ac/sc gene expression to the lateral column in the developing CNS. The concerted action of ftz and odd along the A/P axis, and DER and dpp along the D/V axis, can account for the activation of ac/sc in one eighth of its proneural clusters. Combinations of other patterning genes probably act similarly to initiate AS-C proneural gene expression in the other regions of the developing CNS and PNS.
Scattered throughout the AS-C are regulatory regions that decode the positional information contained within the checkerboard pattern of spatial regulators and translate it into the transcriptional activation of ac, sc, and l'sc. The locations of many of these regulatory regions have been identified through the use of DNA rearrangements within the AS-C and of reporter gene constructs that contain specific genomic regions from the AS-C region (7). For example, ac and sc share a number of regulatory regions located between the two genes that function to activate ac and sc, but not l'sc, in identical patterns throughout development. It is thought that the gene products of many of the patterning genes act directly through these regulatory regions to create the precise and invariant pattern of AS-C-expressing proneural clusters.
3. Resolution of Proneural Cell Clusters: The Singling out of the Neural Precursor
Within each proneural cluster, AS-C expression quickly becomes restricted to one (or a few) cells ) Fig. 1). This cell initiates ase expression, enlarges, and segregates as a neural precursor to a position below the proneural cluster from which it arose. All clusters exhibit identical dynamics of proneural gene expression, which directly reflect the cell-fate decisions made by the cells of a cluster. Initially, all cells express one or more of the proneural genes; proneural gene expression confers on all cells the ability to become a neural precursor. Then, via a cell-cell communication pathway mediated by the Notch signaling pathway, one cell comes to express the proneural genes to the highest level. This cell is the presumptive neural precursor; it then acts, again through the Notch pathway, to inhibit and eventually to extinguish proneural gene expression from all other cells of the cluster (8). The cells that lose proneural gene expression are directed toward the epidermal fate. Thus the fate of cells within a proneural cluster correlates—not with the initial expression of the proneural genes—but rather with the fate of proneural gene expression in that cell: cells that retain proneural gene expression become neural precursors, whereas cells that lose proneural gene expression are directed toward the epidermal fate.
The analysis of the AS-C provides a paradigm for how specific cell fates (and cellular structures ) arise in reproducible and invariant patterns, but many questions still remain with respect to the AS-C. For example, roughly half of all CNS neural precursors still form in embryos devoid of AS-C function. Thus other “proneural genes” must exist that promote neural precursor formation in the CNS. In addition, AS-C function is not restricted to the nervous system. The gene l'sc mediates the initial selection of muscle progenitor cells in the mesoderm in much the same way that it (as well as ac and sc) single out neural precursors in the ectoderm. Furthermore, Galant et al (9) recently demonstrated that a butterfly AS-C homologue is expressed in, and thus may promote the formation of, the precursors to the pigment-producing scale cells that cover the butterfly wing. Thus, AS-C genes may mediate additional developmental events in Drosophila and other animals. Finally, in Drosophila the AS-C is an integral unit composed of four structurally related genes. Has the overall genomic structure of the AS-C and the relative position and expression of each AS-C gene been strongly conserved throughout evolution, as is observed for the genes of the homeobox clusters? Or does the AS-C and its constituent genes display significant plasticity over evolutionary time? Research on the AS-C has led to an understanding of some of the molecular mechanisms that specify particular cell types and those that determine where and when these cell types form. It is likely that further insights will derive from continued analysis of AS-C expression, regulation, and function.
References
1. A. Garcia-Bellido (1979) Genetics 91, 491–520.
2. C. Stern (1954) American Scientist 42, 213–247.
3. S. Campuzano, L. Carramolino, C. V. Cabrera, M. Ruiz-Gomez, R. Villares, A. Boronat, and J. Modolell (1985) Cell 40, 327–338.
4. C. Murre, P. Schonleber-McCaw, H. Vaessin, M. Caudy, L. Y. Jan, Y. N. Jan, C. V. Cabrera, J. N. Buskin, S. D. Hauschka, A. B. Lassar, H. Weintraub, and D. Baltimore (1989) Cell 58, 537–544.
5. J. B. Skeath, G. Panganiban, J. Selegue, and S. B. Carroll (1992) Genes Dev. 6, 2606–2619.
6. J. L. Gomez-Skarmeta, R. Diez del Corral, E. De La Calle-Mustienes, D. Ferres-Marco, and J. Modolell (1996) Cell, 85, 95–105.
7. J. L. Gomez-Skarmeta, I. Rodriguez, C. Martinez, J. Culi, D. Ferres-Marco, D. Beamonte, and J.
Modolell (1995) Genes Dev. 9, 1869–1882.
8. J. A. Campos-Ortega (1993) In The Development of Drosophila melanogaster (M. Bate and A. Martinez-Arias, eds.), Cold Spring Harbor Press, Cold Spring Harbor, NY, pp. 1091–1130.
9. R. Galant, J. B. Skeath, S. Paddock, D. L. Lewis, and S. B. Carroll (1998) Current Biology 8, 807–813.

|
|
|
|
دراسة يابانية لتقليل مخاطر أمراض المواليد منخفضي الوزن
|
|
|
|
|
|
|
اكتشاف أكبر مرجان في العالم قبالة سواحل جزر سليمان
|
|
|
|
|
|
|
المجمع العلمي ينظّم ندوة حوارية حول مفهوم العولمة الرقمية في بابل
|
|
|