النبات

مواضيع عامة في علم النبات

الجذور - السيقان - الأوراق

النباتات الوعائية واللاوعائية

البذور (مغطاة البذور - عاريات البذور)

الطحالب

النباتات الطبية


الحيوان

مواضيع عامة في علم الحيوان

علم التشريح

التنوع الإحيائي

البايلوجيا الخلوية


الأحياء المجهرية

البكتيريا

الفطريات

الطفيليات

الفايروسات


علم الأمراض

الاورام

الامراض الوراثية

الامراض المناعية

الامراض المدارية

اضطرابات الدورة الدموية

مواضيع عامة في علم الامراض

الحشرات


التقانة الإحيائية

مواضيع عامة في التقانة الإحيائية


التقنية الحيوية المكروبية

التقنية الحيوية والميكروبات

الفعاليات الحيوية

وراثة الاحياء المجهرية

تصنيف الاحياء المجهرية

الاحياء المجهرية في الطبيعة

أيض الاجهاد

التقنية الحيوية والبيئة

التقنية الحيوية والطب

التقنية الحيوية والزراعة

التقنية الحيوية والصناعة

التقنية الحيوية والطاقة

البحار والطحالب الصغيرة

عزل البروتين

هندسة الجينات


التقنية الحياتية النانوية

مفاهيم التقنية الحيوية النانوية

التراكيب النانوية والمجاهر المستخدمة في رؤيتها

تصنيع وتخليق المواد النانوية

تطبيقات التقنية النانوية والحيوية النانوية

الرقائق والمتحسسات الحيوية

المصفوفات المجهرية وحاسوب الدنا

اللقاحات

البيئة والتلوث


علم الأجنة

اعضاء التكاثر وتشكل الاعراس

الاخصاب

التشطر

العصيبة وتشكل الجسيدات

تشكل اللواحق الجنينية

تكون المعيدة وظهور الطبقات الجنينية

مقدمة لعلم الاجنة


الأحياء الجزيئي

مواضيع عامة في الاحياء الجزيئي


علم وظائف الأعضاء


الغدد

مواضيع عامة في الغدد

الغدد الصم و هرموناتها

الجسم تحت السريري

الغدة النخامية

الغدة الكظرية

الغدة التناسلية

الغدة الدرقية والجار الدرقية

الغدة البنكرياسية

الغدة الصنوبرية

مواضيع عامة في علم وظائف الاعضاء

الخلية الحيوانية

الجهاز العصبي

أعضاء الحس

الجهاز العضلي

السوائل الجسمية

الجهاز الدوري والليمف

الجهاز التنفسي

الجهاز الهضمي

الجهاز البولي


المضادات الميكروبية

مواضيع عامة في المضادات الميكروبية

مضادات البكتيريا

مضادات الفطريات

مضادات الطفيليات

مضادات الفايروسات

علم الخلية

الوراثة

الأحياء العامة

المناعة

التحليلات المرضية

الكيمياء الحيوية

مواضيع متنوعة أخرى

الانزيمات
Lambda Recombination Occurs in an Intasome
المؤلف:
JOCELYN E. KREBS, ELLIOTT S. GOLDSTEIN and STEPHEN T. KILPATRICK
المصدر:
LEWIN’S GENES XII
الجزء والصفحة:
17-4-2021
3400
Lambda Recombination Occurs in an Intasome
KEY CONCEPTS
- Lambda integration takes place in a large complex that also includes the host protein IHF.
- The excision reaction requires Int and Xis and recognizes the ends of the prophage DNA as substrates.
Unlike the Cre/lox recombination system, which requires only the enzyme and the two recombining sites, phage lambda recombination occurs in a large structure and has different components for each direction of the reaction (integration versus excision).
The host protein IHF is required for both integration and excision. IHF is a 20-kD protein of two different subunits, which are encoded by the genes himA and himD. IHF is not an essential protein in E. coli and is not required for homologous bacterial recombination. It is one of several proteins with the ability to wrap DNA on a surface.
Mutations in the him genes prevent lambda site–specific recombination and can be suppressed by mutations in λint, which suggests that IHF and Int interact. Site-specific recombination can
be performed in vitro by Int and IHF.
The in vitro reaction requires supercoiling in attP, but not in attB. When the reaction is performed in vitro between two supercoiled DNA molecules, almost all of the supercoiling is retained by the products. Thus, there cannot be any free intermediates in which strand rotation could occur. This was one of the early hints that the reaction proceeds through a Holliday junction. We now know that the reaction proceeds by the mechanism typical of this class of enzymes, which is related to the topoisomerase I mechanism .
Int has two different modes of binding. The C-terminal domain behaves like the Cre recombinase. It binds to inverted sites at the core sequence, positioning itself to make the cleavage and ligation reactions on each strand at the positions illustrated in FIGURE 1. The N-terminal domain binds to sites in the arms of attP that have a different consensus sequence. This binding is responsible for the aggregation of subunits into the intasome. The two domains probably bind DNA simultaneously, thus bringing the arms of attP close to the core.
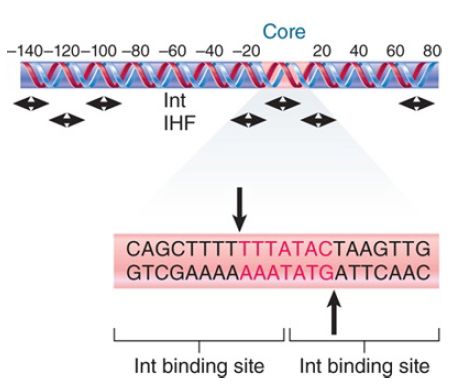
FIGURE 1. Int and IHF bind to different sites in attP. The Int recognition sequences in the core region include the sites of cutting.
IHF binds to sequences of about 20 bp in attP. The IHF-binding sites are approximately adjacent to sites where Int binds. Xis binds to two sites located close to one another in attP, so that the protected region extends over 30 to 40 bp. Together, Int, Xis, and IHF cover virtually all of attP. The binding of Xis changes the organization of the DNA so that it becomes inert as a substrate for the integration reaction.
When Int and IHF bind to attP, they generate a complex in which all the binding sites are pulled together on the surface of a protein. Supercoiling of attP is needed for the formation of this intasome. The only binding sites in attB are the two Int sites in the core. Int does not bind directly to attB in the form of free DNA, though. The intasome is the intermediate that “captures” attB, as indicated schematically in FIGURE 2.
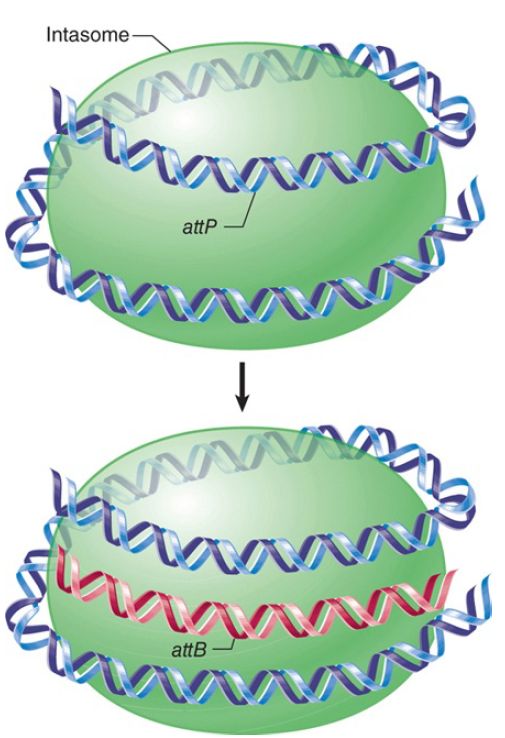
FIGURE 2. Multiple copies of Int protein may organize attP into an intasome, which initiates site-specific recombination by recognizing attB on free DNA.
According to this model, the initial recognition between attP and attB does not depend directly on DNA homology, but instead is determined by the ability of Int proteins to recognize both att
sequences. The two att sites then are brought together in an orientation predetermined by the structure of the intasome. Sequence homology becomes important at this stage, when it is required for the strand-exchange reaction.
The asymmetry of the integration and excision reactions is shown by the fact that Int can form a similar complex with attR only if Xis is added. This complex can pair with a condensed complex that Int forms at attL. IHF is not needed for this reaction. A significant difference between lambda integration/excision and the recombination reactions catalyzed by Cre or Flp is that Intcatalyzed reactions bind the regulatory sequences in the arms of the target sites, bending the DNA and allowing interactions between arm and core sites that drive each reaction to its conclusion. This is why each lambda reaction is irreversible, whereas recombination catalyzed by Cre or Flp is reversible.
Crystal structures of λ-Int tetramers show that, like other recombinases, the tetramer has two active and two inactive subunits that switch roles during recombination. Allosteric interactions triggered by arm-binding control structural transitions in the tetramer that drive the reaction.
Much of the complexity of site-specific recombination may be caused by the need to regulate the reaction so that integration occurs preferentially when the virus is entering the lysogenic state, whereas excision is preferred when the prophage is entering the lytic cycle. By controlling the amounts of Int and Xis, the appropriate reaction will occur.
 الاكثر قراءة في مواضيع عامة في الاحياء الجزيئي
الاكثر قراءة في مواضيع عامة في الاحياء الجزيئي
 اخر الاخبار
اخر الاخبار
اخبار العتبة العباسية المقدسة

الآخبار الصحية















 قسم الشؤون الفكرية يصدر كتاباً يوثق تاريخ السدانة في العتبة العباسية المقدسة
قسم الشؤون الفكرية يصدر كتاباً يوثق تاريخ السدانة في العتبة العباسية المقدسة "المهمة".. إصدار قصصي يوثّق القصص الفائزة في مسابقة فتوى الدفاع المقدسة للقصة القصيرة
"المهمة".. إصدار قصصي يوثّق القصص الفائزة في مسابقة فتوى الدفاع المقدسة للقصة القصيرة (نوافذ).. إصدار أدبي يوثق القصص الفائزة في مسابقة الإمام العسكري (عليه السلام)
(نوافذ).. إصدار أدبي يوثق القصص الفائزة في مسابقة الإمام العسكري (عليه السلام)