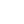النبات

مواضيع عامة في علم النبات

الجذور - السيقان - الأوراق

النباتات الوعائية واللاوعائية

البذور (مغطاة البذور - عاريات البذور)

الطحالب

النباتات الطبية


الحيوان

مواضيع عامة في علم الحيوان

علم التشريح

التنوع الإحيائي

البايلوجيا الخلوية


الأحياء المجهرية

البكتيريا

الفطريات

الطفيليات

الفايروسات


علم الأمراض

الاورام

الامراض الوراثية

الامراض المناعية

الامراض المدارية

اضطرابات الدورة الدموية

مواضيع عامة في علم الامراض

الحشرات


التقانة الإحيائية

مواضيع عامة في التقانة الإحيائية


التقنية الحيوية المكروبية

التقنية الحيوية والميكروبات

الفعاليات الحيوية

وراثة الاحياء المجهرية

تصنيف الاحياء المجهرية

الاحياء المجهرية في الطبيعة

أيض الاجهاد

التقنية الحيوية والبيئة

التقنية الحيوية والطب

التقنية الحيوية والزراعة

التقنية الحيوية والصناعة

التقنية الحيوية والطاقة

البحار والطحالب الصغيرة

عزل البروتين

هندسة الجينات


التقنية الحياتية النانوية

مفاهيم التقنية الحيوية النانوية

التراكيب النانوية والمجاهر المستخدمة في رؤيتها

تصنيع وتخليق المواد النانوية

تطبيقات التقنية النانوية والحيوية النانوية

الرقائق والمتحسسات الحيوية

المصفوفات المجهرية وحاسوب الدنا

اللقاحات

البيئة والتلوث


علم الأجنة

اعضاء التكاثر وتشكل الاعراس

الاخصاب

التشطر

العصيبة وتشكل الجسيدات

تشكل اللواحق الجنينية

تكون المعيدة وظهور الطبقات الجنينية

مقدمة لعلم الاجنة


الأحياء الجزيئي

مواضيع عامة في الاحياء الجزيئي


علم وظائف الأعضاء


الغدد

مواضيع عامة في الغدد

الغدد الصم و هرموناتها

الجسم تحت السريري

الغدة النخامية

الغدة الكظرية

الغدة التناسلية

الغدة الدرقية والجار الدرقية

الغدة البنكرياسية

الغدة الصنوبرية

مواضيع عامة في علم وظائف الاعضاء

الخلية الحيوانية

الجهاز العصبي

أعضاء الحس

الجهاز العضلي

السوائل الجسمية

الجهاز الدوري والليمف

الجهاز التنفسي

الجهاز الهضمي

الجهاز البولي


المضادات الميكروبية

مواضيع عامة في المضادات الميكروبية

مضادات البكتيريا

مضادات الفطريات

مضادات الطفيليات

مضادات الفايروسات

علم الخلية

الوراثة

الأحياء العامة

المناعة

التحليلات المرضية

الكيمياء الحيوية

مواضيع متنوعة أخرى

الانزيمات
Electrophoresis of Nucleic Acids
المؤلف:
Wilson, K., Hofmann, A., Walker, J. M., & Clokie, S. (Eds.)
المصدر:
Wilson and Walkers Principles and Techniques of Biochemistry and Molecular Biology
الجزء والصفحة:
8th E , P240-246
2026-04-01
57
Agarose Gel Electrophoresis of DNA
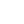
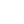
For the majority of DNA samples, electrophoretic separation is carried out in aga rose gels. This is because most DNA molecules and their fragments that are analysed routinely are considerably larger than proteins and since such fragments would be unable to enter a polyacrylamide gel, the larger pore size of an agarose gel is required. For example, the commonly used plasmid pBR322 has an M r of 2.4 × 106. However, rather than using such large numbers, it is more convenient to refer to DNA size in terms of the number of base-pairs. Although, originally, DNA size was referred to in terms of base-pairs ( bp) or kilobase-pairs (kbp), it has now become the accepted nomenclature to abbreviate kbp to simply kb when referring to double-stranded DNA. In that nomenclature, pBR322 has a size of 4.36 kb; note that even a small restriction fragment of 1 kb has an M r of 620000. When talking about single-stranded DNA it is common to refer to size in terms of nucleotides ( nt). Since the charge per unit length (owing to the phosphate groups) in any given fragment of DNA is the same, all DNA samples should move towards the anode with the same mobility under an applied electrical fi eld. However, separation in agarose gels is achieved because of resistance to their movement caused by the gel matrix. The largest molecules will have the most difficulty passing through the gel pores (very large molecules may even be blocked completely), whereas the smallest molecules will be relatively unhindered. Consequently, the mobility of DNA molecules during gel electrophoresis will depend on size, the smallest molecules moving fastest. This is analogous to the separation of proteins in SDS–polyacrylamide gels, although the analogy is not perfect, as double-stranded DNA molecules form relatively stiff rods and while it is not completely understood how they pass through the gel, it is probable that long DNA molecules pass through the gel pores end-on. While passing through the pores, a DNA molecule will experience drag; so the longer the molecule, the more it will be retarded by each pore. Sideways movement may become more important for very small double-stranded DNA and for the more flexible single-stranded DNA. It will be obvious from the above that gel concentrations must be chosen to suit the size range of the molecules to be separated. Gels containing 0.3% agarose will separate double-stranded DNA molecules of between 5 and 60 kb in size, whereas 2% gels are used for samples of between 0.1 and 3 kb. Many laboratories routinely use 0.8% gels, which are suitable for separating DNA molecules in the range 0.5–10 kb. Since agarose gels separate DNA according to size, the M r of a DNA fragment may be determined from its electrophoretic mobility by running a number of standard DNA markers of known M r on the same gel. This is most conveniently achieved by running a sample of bacteriophage λ DNA (49 kb) that has been cleaved with a restriction enzyme such as Eco RI. Since the base sequence of λ DNA is known, and the cleavage sites for Eco RI are known, this generates fragments of accurately known size (Figure 1).
Fig1. Photograph showing four tracks from a 0.8% agarose submarine gel. The gel was run at 40 V in TRIS/borate/EDTA buffer for 16 h, stained with ethidium bromide and viewed under ultraviolet light. Sample loadings were about 0.5 µg of DNA per track. Tracks 1 and 2, λ DNA (49 kb). Track 3, λ DNA cleaved with the enzyme Eco RI to generate fragments of the following size (in order from the origin): 21.80 kb, 7.52 kb, 5.93 kb, 5.54 kb, 4.80 kb, 3.41 kb. Track 4, λ DNA cleaved with the enzyme Hind III to generate fragments of the following size (in order from the origin): 23.70 kb, 9.46 kb, 6.75 kb, 4.26 kb, 2.26 kb, 1.98 kb. (Courtesy of Stephen Boffey, University of Hertfordshire.)
DNA gels are invariably run as horizontal, submarine or submerged gels; so named because such a gel is totally immersed in buffer. Agarose, dissolved in gel buffer by boiling, is poured onto a glass or plastic plate, surrounded by a wall of adhesive tape or a plastic frame to provide a gel about 3 mm in depth. Loading wells are formed by placing a plastic well-forming template or comb in the poured gel solution, and removing this comb once the gel has set. The gel is placed in the electrophoresis tank, covered with buffer, and samples loaded by directly injecting the sample into the wells. Samples are prepared by dissolving them in a buffer solution that contains sucrose, glycerol or Ficoll ® , which makes the solution dense and allows it to sink to the bottom of the well. A dye such as bromophenol blue is also included in the sample solvent; it makes it easier to see the sample that is being loaded and also acts as a marker of the electrophoresis front. No stacking gel is needed for the electrophoresis of DNA, because the mobilities of DNA molecules are much greater in the well than in the gel, and therefore all the molecules in the well pile up against the gel within a few minutes of the current being turned on, forming a tight band at the start of the run. General purpose gels are approximately 25 cm long and 12 cm wide, and are run at a voltage gradient of about 1.5 V cm−1 overnight. A higher voltage would cause excessive heating. For rapid analyses that do not need extensive separation of DNA molecules, it is common to use minigels that are less than 10 cm long. In this way information can be obtained in 2–3 h.
Once the system has been run, the DNA in the gel needs to be stained and visualized. The reagent most widely used include the fluorescent dyes ethidium bromide and SYBR ® Green. The gel is rinsed gently in a solution of the fluorescent dye and then viewed under ultraviolet light (300 nm wavelength); alternatively, the dye can be dissolved in liquefied agarose just before pouring the gel. Dyes such as ethidium bromide and SYBR ® Green are molecules that bind between the stacked base-pairs of DNA (i.e. they intercalate). The dye concentration therefore builds up at the site of the DNA bands and under ultraviolet light, the DNA bands fluoresce orange-red (ethidium bromide) or green (SYBR ® Green). As little as 10 ng of DNA can be visualised as a 1 cm wide band. It should be noted that extensive viewing of the DNA with ultraviolet light can result in damage of the DNA by nicking and base-pair dimerisation. This is of no consequence if a gel is only to be viewed, but obviously viewing of the gel should be kept to a minimum if the DNA is to be recovered. It is essential to protect one’s eyes by wearing goggles when ultraviolet light is used. If viewing of gels under ultraviolet is carried out for long periods, a plastic mask that covers the whole face should be used to avoid ‘sunburn’.
DNA Sequencing Gels
DNA sequencing gels have been a ‘workhorse’ technique for the molecular biologist, but have now been replaced by automated methods such as dideoxy sequencing for routine applications. However, for some particular applications, such as DNA footprinting, sequencing gels are still used.
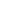
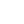
Whereas agarose gel electrophoresis of DNA is highly suitable for relatively short DNA molecules, a different form of electrophoresis has to be used when DNA sequences are to be determined. Whichever DNA sequencing method is used, the final analysis usually involves separating single-stranded DNA molecules shorter than about 1000 nt and differing in size by only 1 nt. To achieve this, it is necessary to have a small-pored gel and so acrylamide gels are used instead of agarose. For example, 3.5% polyacrylamide gels are used to separate DNA in the range 80–1000 nt and 12% gels to resolve fragments of between 20 and 100 nt. If a wide range of sizes needs to be analysed, it is often convenient to run a gradient gel, for example from 3.5% to 7.5%. Sequencing gels are run in the presence of denaturing agents, urea and formamide. Since it is necessary to separate DNA molecules that are very similar in size, DNA sequencing gels tend to be very long (100 cm) to maximise the separation achieved. A typical DNA sequencing gel is shown in Figure 2.
Fig2. Autoradiograph of a DNA sequencing gel. Samples were prepared using the Sanger dideoxy method of DNA sequencing. Each set of four samples was loaded into adjacent tracks, indicated by A,C, G and T, depending on the identity of the dideoxyribonucleotide used for that sample. Two sets of samples were labelled with 35 S (1 and 3) and one was labelled with 32 P (2). It is evident that 32 P generates darker, but more diffuse bands than does 35 S, making the bands nearer the bottom of the autoradiograph easy to see. However, the broad bands produced by 32 P cannot be resolved near the top of the autoradiograph, making it impossible to read a sequence from this region. The much sharper bands produced by 35 S allow sequences to be read with confidence along most of the autoradiograph and so a longer sequence of DNA can be obtained from a single gel.
As mentioned above, electrophoresis in agarose can be used as a preparative method for DNA. The DNA bands of interest can be cut out of the gel and the DNA recovered by: (a) electroelution, (b) macerating the gel piece in buffer, centrifuging and collecting the supernatant or (c), if low melting point agarose is used, melting the gel piece and diluting with buffer. In each case, the DNA is finally recovered by precipitation of the supernatant with ethanol.
Pulsed-Field Gel Electrophoresis
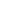
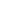
The agarose gel methods for DNA described above can fractionate DNA of 60 kb or less. The introduction of pulsed-fi eld gel electrophoresis (PFGE) and the further development of variations on the basic technique means that nowadays DNA fragments up to 2 × 103 kb can be separated. Essentially, this allows the separation of whole chromosomes by electrophoresis. The method basically involves electrophoresis in agarose, where two electric fi elds are applied alternately at different angles for defined time periods (e.g. 60 s). Activation of the first electric fi eld causes the coiled molecules to be stretched in the horizontal plane and start to move through the gel. Interruption of this fi eld and application of the second fi eld force the molecule to move in the new direction. Owing to a length-dependent relaxation behaviour when a long-chain molecule undergoes conformational change in an electric fi eld, the smaller a molecule, the quicker it realigns itself with the new fi eld and is able to continue moving through the gel. Larger molecules take longer to realign. In this way, with continual reversing of the field, smaller molecules draw ahead of larger molecules and separate according to size. PFGE has proved particularly useful in identifying the course of outbreaks of bacterial food-borne illness (e.g. Salmonella infections). Having isolated the bacterial pathogen responsible for the illness from an individual, the DNA is isolated and cleaved into large fragments, which are separated by PFGE. For example, DNA from Salmonella species, when digested with the restriction enzyme Xba I, gives around 15 fragments ranging from 25 kb to 680 kb. This pattern of fragments, or ‘ fingerprint’, is unique to that strain. If the same fingerprint is found from bacteria from other infected people, then it can be assumed that they were all infected from a common source. Thus, by comparing their eating habits, food sources, etc. the source of the infection can be traced to a restaurant, food item, etc. Figure 3 shows the restriction patterns from different strains of Neisseria meningitidis.
Fig3. PFGE separation of the digestion pattern produced with the restriction enzyme Nhe I, of 21 strains of Neisseria meningitidis . There are two molecular-weight marker tracks at either end of the gel. (Courtesy of Dr Giovanna Morelli, Max Planck Institute for Molecular Genetics, Berlin, Germany.)
Electrophoresis of RNA
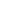
Like that of DNA, electrophoresis of RNA is usually carried out in agarose gels, and the principle of the separation, based on size, is the same. Often, one requires a rapid method for checking the integrity of RNA immediately following extraction and before deciding whether to process it further. This can be achieved easily by electrophoresis in a 2% agarose gel in about 1 h. Ribosomal RNAs (18 S and 28 S) are clearly resolved and any degradation (seen as a smear) or DNA contamination is seen easily. This can be achieved on a 2.5–5% acrylamide gradient gel with an overnight run. Both these methods involve running native RNA. There will almost certainly be some secondary structure within the RNA molecule owing to intramolecular hydro gen bonding (see, for example, the clover leaf structure of tRNA). For this reason, native RNA run on gels can be stained and visualised with ethidium bromide. However, if the study objective is to determine RNA size by gel electrophoresis, then full denaturation of the RNA is needed to prevent hydrogen bond formation within or even between polynucleotides that will otherwise affect the electrophoretic mobility. There are three denaturing agents (formaldehyde, glyoxal and methylmercuric hydroxide) that are compatible with both RNA and agarose. Either one of these may be incorporated into the agarose gel and electrophoresis buffer, and the sample is heat denatured in the presence of the denaturant prior to electrophoresis. After heat denaturation, each of these agents forms adducts with the amino groups of guanine and uracil, thereby preventing hydrogen bond reformation at room temperature during electrophoresis. It is also necessary to run denaturing gels if the RNA is to be blotted (Northern blots) and probed, to ensure that the base sequence is available to the probe. Denatured RNA stains only very weakly with ethidium bromide, so acridine orange is commonly used to visualise RNA on denaturing gels. However, it should be noted that many workers will be using radiolabelled RNA and accordingly identify bands by autoradiography. An example of the electrophoresis of RNA is shown in Figure4.
Fig4. Separation of yeast RNA on a 1.5% agarose gel. (Courtesy of Dr Tomas Masek, Department of Genetics and Microbiology, Charles University, Prague, Czech Republic.)
 الاكثر قراءة في مواضيع عامة في الاحياء الجزيئي
الاكثر قراءة في مواضيع عامة في الاحياء الجزيئي
 اخر الاخبار
اخر الاخبار
اخبار العتبة العباسية المقدسة

الآخبار الصحية















 قسم الشؤون الفكرية يصدر كتاباً يوثق تاريخ السدانة في العتبة العباسية المقدسة
قسم الشؤون الفكرية يصدر كتاباً يوثق تاريخ السدانة في العتبة العباسية المقدسة "المهمة".. إصدار قصصي يوثّق القصص الفائزة في مسابقة فتوى الدفاع المقدسة للقصة القصيرة
"المهمة".. إصدار قصصي يوثّق القصص الفائزة في مسابقة فتوى الدفاع المقدسة للقصة القصيرة (نوافذ).. إصدار أدبي يوثق القصص الفائزة في مسابقة الإمام العسكري (عليه السلام)
(نوافذ).. إصدار أدبي يوثق القصص الفائزة في مسابقة الإمام العسكري (عليه السلام)